Decluttering ordination plots in vegan part 2: orditorp()
In the earlier post in this series I looked at the ordilabel() function to help tidy up ordination biplots in vegan. An alternative function vegan provides is orditorp(), the last four letters abbreviating the words text or points. That is a pretty good description of what orditorp() does; it draws sample or species labels using text where there is room and where there isn’t a plotting character is drawn instead. Essentially it boils down to being a one stop shop for calls to text() or points() as needed. Let’s see how it works…
As with last time out, I’ll illustrate how orditorp() works via a PCA biplot for the Dutch dune meadow data.
## load vegan and the data
require(vegan)
data(dune)
ord <- rda(dune) # PCA of Dune data
## species priority; which species drawn last, i.e. on top
priSpp <- diversity(dune, index = "invsimpson", MARGIN = 2)
## sample priority
priSite <- diversity(dune, index = "invsimpson", MARGIN = 1)
## scaling to use
scl <- 3I won’t explain any of the code above; it is the same as that used in the earlier post where an explanation was also provided. orditorp() takes an ordination object as the first argument and in addition the display argument controls which set of scores is displayed. Note that orditorp() can only plot one set of scores at a time, which as we’ll see in a minute is not exactly ideal nor foolproof. Like ordilabel(), you are free to specify the importance of each sample or species via argument priority. In ordilable() the priority controlled the plotting order such that those samples or species with high priority were plotted last (uppermost). Instead, orditorp() draws labels for samples or species (if it can) for those with the highest priority first.
So we have something to talk to, recreate the basic samples and species biplot as used in the previous post but updated to use orditorp()
plot(ord, type = "n", scaling = 3)
orditorp(ord, display = "sites", priority = priSite, scaling = scl,
col = "blue", cex = 1, pch = 19)
## You may prefer separate plots, but here species as well
orditorp(ord, display = "species", priority = priSpp, scaling = scl,
col = "forestgreen", pch = 2, cex = 1)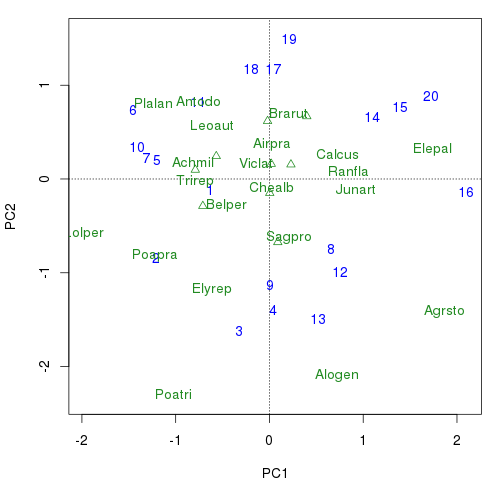
orditorp()The behaviour or orditorp() should now be reasonably clear; labels are drawn for sample or species only if there is room to do so, with a point being used instead. orditorp() isn’t perfect by any means. Because it can only drawn one set of scores at a time, there is no easy way to stop the species labels plotting over the sample labels and vice versa.
How it works is, first orditorp() calculates the heights and widths of the labels, adds a bit of space to this (more on this later) and then works out if the box given by the current sample or species label width/height, centred on the axis score coordinate, will obscure the label boxes of any labels previously drawn. If the label box doesn’t obscure any previous label boxes the label is drawn at the sample or species score coordinates. If it does obscure an existing label then a point is drawn instead. orditorp() draws the labels in order of priority and as it draws each subsequent label it checks to see if previous labels are not obscured.
This process isn’t infallible of course; for example the second highest priority sample or species could lie very close to the highest priority one in ordination space and if so orditorp() would not draw a label for this second highest priority sample or species because it would obscure the label of the highest priority one.
The amount of spacing or padding around each label is specified via the air argument which has a default of 1. air is interpreted as the proportion of half the label width or height that the label occupies. The default of 1 therefore means that in fact there is no additional spacing beyond the confines of the box that encloses the label. If air is greater than 1 proportionally more padding is added whilst values less than 1 indicate that labels can overlap. The figure below shows the species scores only with two values for air. In the left hand panel air = 2 is used and the labels are padded either side of the label by the entire string width or height. The right hand panel uses air = 0.5 which allows labels to overlap by up to a quarter of the string width or height in any direction from the plotting coordinate (in other words, the box that cannot be obscured when plotting subsequent labels is half the string width wide and half the string height high, centred on the plotting coordinates for the label).
layout(matrix(1:2, ncol = 2))
op <- par(mar = c(5,4,4,1) + 0.1)
## site/sample scores
plot(ord, type = "n", scaling = 3, main = expression(air == 2), cex = 1)
orditorp(ord, display = "species", priority = priSite, scaling = scl,
col = "forestgreen", cex = 1, pch = 2, air = 2)
## Species scores
plot(ord, type = "n", scaling = 3, main = expression(air == 0.5), cex = 1)
orditorp(ord, display = "species", priority = priSpp, scaling = scl,
col = "forestgreen", pch = 2, cex = 1, air = 0.5)
par(op)
layout(1)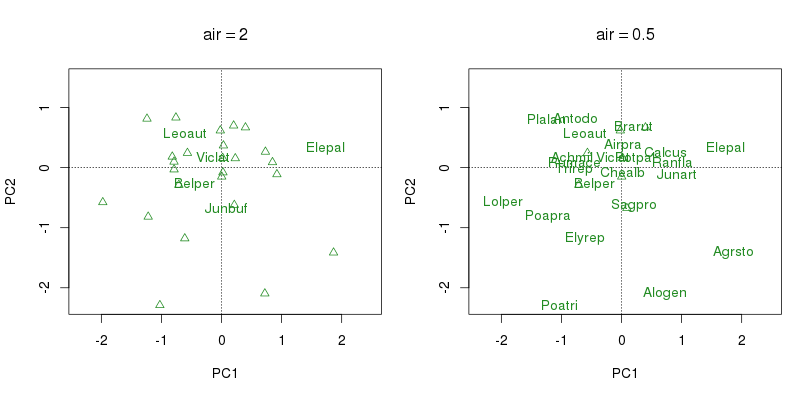
orditorp() showing the effect of changing argument air.One point that should be noted is that orditorp() doesn’t stop labels and points from overlaying one another, though as the labels are drawn after the points they shouldn’t get obscured too much. We could improve the situation a bit by drawing an opaque box around the label, or even make it partially transparent, so that the label always stood out from the plotting points. Although we’d run the risk of hiding points under labels and thus hiding information from the person looking at the figure.
One additional point to make is that orditorp() returns a logical vector indicating which sample or species scores were drawn with labels (TRUE) or points (FALSE), which might be useful for further plotting or adding to the diagram. So there were have orditorp().
Next time I’ll take a look at ordipointlabel() which tackles the problem of producing a tidy ordination diagram in a far more complex way than either ordilabel() or orditorp().

